

312 Church St. SE
Minneapolis, MN 55455
United States
John D.
Lipscomb
We investigate the mechanisms of oxygenases, the enzymes nature uses to catalyze the incorporation of oxygen into biological molecules. Oxygen incorporation reactions are important for a wide variety of reasons, ranging from activation of steroid hormones in humans to the detoxification of compounds in the environment. The oxygenases we study contain a metal which is the key to the chemistry they catalyze.
We investigate the mechanisms of oxygenases, the enzymes nature uses to catalyze the incorporation of oxygen into biological molecules. Oxygen incorporation reactions are important for a wide variety of reasons, ranging from activation of steroid hormones in humans to the detoxification of compounds in the environment. The oxygenases we study contain a metal which is the key to the chemistry they catalyze. Enzyme mechanisms involve both the chemical reactions occurring at the active site and the regulation of the reaction imposed by the complex protein structure. Consequently, we use many types of biochemical and physical techniques including transient kinetics, site directed mutagenesis, diagnostic substrate reactions, and EPR spectroscopy. This is extended to X-ray crystallography and other spectroscopies through collaborations.
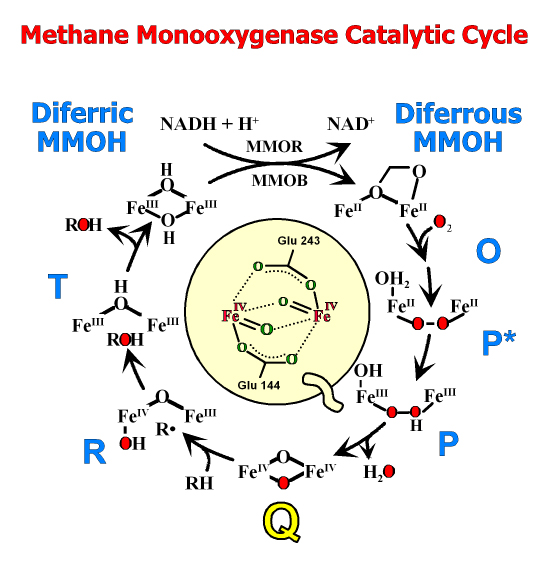
Currently, we are studying two large families of oxygenases. The first is typified by methane monooxygenases, the enzyme primarily responsible for preventing the vast amounts of biologically generated methane from reaching the atmosphere. This enzyme splits O2 and incorporates one atom of oxygen into methane to form methanol, while reducing the second atom to water. Another enzyme we have found from this family is essential for the biosynthesis of many important antibiotics and chemotherapy drugs. The second oxygenase family includes several dioxygenase enzymes that also split O2 but incorporate both atoms into biomolecules. These enzymes are the major means by which the enormous quantities of aromatic compounds that enter the environment annually are reassimilated into the carbon cycle. Recently, we have developed methods to trap reaction cycle intermediates of these enzymes in single crystals for direct structural analysis. Related collaborative projects include studies of the biosynthesis of biofuels, penicillin, fosfomycin, and ethylene (a plant hormone).
Jones, J. C., Banerjee, R., Semonis, M. M., Shi, K., Aihara, H., and Lipscomb, J. D. “X-ray Crystal Structures of Methane Monooxygenase Hydroxylase Complexes with Variants of Its Regulatory Component: Correlations with Altered Reaction Cycle Dynamics” Biochemistry, 61, 21-33 (2022). PMC8727504 PubMed
Jacobs, A. J., Banerjee, R., Deweese, D. E., Augustin, B., Babicz, Jr., J. T., Gee, L. B., Sutherlin, K. D., Böttger, L. H., Yoda, Y., Saito, M., Kitao, S., Kobayashi, Y., Seto, M., Tamasaku, K., Lipscomb, J. D., Kiyoung Park, K., and Solomon, E. I. “Nuclear Resonance Vibrational Spectroscopic Definition of the Fe(IV)2 Intermediate Q in Methane Monooxygenase and Its Reactivity” J. Am. Chem. Soc. 143, 16007-16029 (2021) PMC8631202 PubMed
Jones, J. C., Banerjee, R., Shi, K., Semonis, M. M., Aihara, H., Pomerantz, W. C. K., and Lipscomb, J. D. “Soluble Methane Monooxygenase Component Interactions Monitored by 19F NMR”, Biochemistry, 60, 1995-2010 (2021) PMC8345336 PubMed
Banerjee, R. and Lipscomb, J. D. “Small Molecule Tunnels in Metalloenzymes Viewed as Extensions of the Active Site” Acc. Chem. Res., 54, 2185-2195 (2021) PMC8130187 PubMed
Kotandeniya, D., Rogers, M. S., Fernandez, J., Kanugula, S., Hudson, R. H. E., Rodriguez, F., Lipscomb, J. D., and Tretyakova, N. “6-Phenylpyrrolocytidine as a Fluorescent Probe to Examine Nucleotide Flipping Catalyzed by a DNA Repair Protein” Biopolymers, 112, e23405 (2021). doi: 10.1002/bip.23405. PMC7856187 PubMed
Jones, J. C., Banerjee, R., Shi, K., Aihara, H., and Lipscomb, J. D. “Structural Studies of Methylosinus trichosporium OB3b Soluble Methane Monooxygenase Hydroxylase and Regulatory Component Complex Reveal a Transient Substrate Tunnel” Biochemistry, 59, 2946-2961 (2020) PMC7457393 PubMed
Srinivas, V., Banerjee, R., Lebrette, H., Jones, J. C. ………V. K., Yano, J., Lipscomb, J. D., Kern, J. F., and Högbom, M. “High Resolution XFEL Structure of the Soluble Methane Monooxygenase Hydroxylase Complex with its Regulatory Component at Ambient Temperature in Two Oxidation States” J. Am. Chem. Soc. 142, 14249-14266 (2020) PMC7457426 PubMed
Rogers, M. S., and Lipscomb, John D. “Salicylate 5-Hydroxylase: Intermediates in Aromatic Hydroxylation by a Rieske Monooxygenase” Biochemistry, 58, 5305-5319 (2019) PMC6856394 PubMed
Banerjee, R, Jones, J. C. and Lipscomb. J. D. "Soluble Methane Monooxygenase" Ann. Rev. Biochem., 88, 409-431 (2019) PMID: 30633550 PubMed
Sutherlin, K. D., Wasada-Tsutsui, Y., Mbughuni, M. M., Rogers, M. S., Park, K., Lei V. Liu, L. V., Kwak, Y., Srnec, M., Böttger, L. H., Frenette, M., Yoda, Y., Kobayashi, Y., Kurokuzu, M., Saito, M., Seto, M., Hu, M., Zhao, J., Alp, E. E., Lipscomb, J. D., and Solomon, E. I. “NRVS Definition of O2 Intermediates in an Extradiol Dioxygenase: Correlation to Crystallography and Reactivity” J. Am. Chem. Soc., 140, 16495–16513 (2018) PMC6470009 PubMed
Cutsail III, G. E., Banerjee, R., Zhou, A., Que, Jr., L., Lipscomb, J. D., and DeBeer, S. “High-Resolution EXAFS Provides Evidence for a Longer Fe•••Fe Distance in the Q Intermediate of Methane Monooxygenase” J. Am. Chem. Soc., 140, 16807–16820 (2018) PMC6470014 PubMed
Sutherlin, K. D., Rivard, B. S., Böttger, L. H., Liu, L. V., Rogers, M. S., Srnec, M., Park, K., Yoda, Y., Kitao, S., Kobayashi, Y., Saito, M., Seto, M., Hu, M., Zhao, J., Lipscomb, J. D., Solomon, E. I. “NRVS Studies of the Peroxide Shunt Intermediate in a Rieske Dioxygenase and its Relation to the Native FeII O2 Reaction” J. Amer. Chem Soc., 140, 5544-5559 (2018) PMC5973823 PubMed
Komor, A. J., Jasniewski, A. J., Que, L., Jr. and Lipscomb, J. D. “Diiron Monooxygenases in Natural Product Biosynthesis” Nat. Prod. Rep., 35, 646-659 (2018) PMC6051903 PubMed
Castillo, R. G., Banerjee, R., Allpress, C. J., Rohde, G. T., Bill, E., Que, L., Jr., Lipscomb, J. D., and DeBeer, S. “High-Energy Resolution Fluorescence Detected X-ray Absorption of the Q Intermediate of Soluble Methane Monooxygenase” J. Amer. Chem Soc., 139, 18024-18033 (2017) PMC5729100 PubMed
Oloo, W., Banerjee, R., Lipscomb, J. D., and Que, L., Jr. "Equilibrating (L)FeIII–OOAc and (L)FeV (O) Species in Hydrocarbon Oxidations by Bio-Inspired Nonheme Iron Catalysts using H2O2 and AcOH" J. Amer. Chem Soc., 139, 17313-17326 (2017) PMC5768304 PubMed
Banerjee, R., Komor, A. J., and Lipscomb, J. D. “Use of Isotopes and Isotope Effects for Investigations of Diiron Oxygenase Mechanisms” Meth. Enzymol. 596, 239-290 (2017) PubMed
Komor, A. J., Rivard, B. S., Fan, R., Guo, Y., Que, L., Jr., and Lipscomb, J. D. “CmlI N-Oxygenase Catalyzes the Final Three Steps in Chloramphenicol Biosynthesis without Dissociation of Intermediates” Biochemistry, 56, 4940-4950 (2017) PMC 5605456 PubMed
Jasniewski, A. J., Komor, A. J., Lipscomb, J. D., and Que, L., Jr. “An Unprecedented (μ-1,1-Peroxo)diferric Structure for the Ambiphilic Orange Peroxo Intermediate of the Nonheme N-Oxygenase CmlI” J. Amer. Chem Soc., 139, 10472–10485 (2017) PMC 5568637 PubMed
Eiden, C., Maize, K. M., Finzel, B. C., Lipscomb, J. D., and Aldrich, C. C. “Rational Optimization of Mechanism-Based Inhibitors through Determination of the Microscopic Rate Constants of Inactivation” J. Amer. Chem. Soc., 139, 7132-7135 (2017) PMC 5590675 PubMed
Komor, A. J., Rivard, B. S., Fan, R., Guo, Y., Que, L., Jr., Lipscomb, J. D. “Mechanism for Six-Electron Aryl-N-Oxygenation by the Non-heme Diiron Enzyme CmlI” J. Am. Chem. Soc., 138, 7411-7421 (2016) PMC
Knoot, C. J., Kovaleva, E. G., Lipscomb, J. D. “Crystal Structure of CmlI, the Arylamine Oxygenase from the Chloramphenicol Biosynthetic Pathway” J. Biol. Inorg. Chem., 21, 589-603 (2016). PMC
Meier, K. K., Rogers, M. S., Kovaleva, E. G.; Mbughuni, M. M.; Bominaar, E. L., Lipscomb, J. D.; and Münck, E. “A Long-lived FeIII-(Hydroperoxo) Intermediate in the Active H200C Variant of Homoprotocatechuate 2,3-Dioxygenase: Characterization by Mössbauer, EPR, and DFT Methods” Inorg. Chem., 54, 10269-10280 (2015) PMC
Kovaleva, E. G., Rogers, M. S., and Lipscomb, J. D. “Structural Basis for Substrate and Oxygen Activation in Homoprotocatechuate 2,3-Dioxygenase: Roles of Conserved Active Site Histidine-200” Biochemistry, 54, 5329-5339 (2015) PMC
Rivard, B. S., Rogers, M. S., Marell, D. J., Neibergall, M. B., Chakrabarty, S.,
Cramer, C. J., and Lipscomb, J. D. “Rate-determining Attack on Substrate Precedes Rieske Cluster Oxidation during cis-Dihydroxylation by Benzoate Dioxygenase” Biochemistry, 54, 4652-4664 (2015) PMC
Banerjee, R., Proshlyakov, Y., Lipscomb, J.D., and Proshlyakov, D.A. “Structure of the key species in the enzymatic oxidation of methane to methanol” Nature, 518(7539), 431-434 (2015) PMC
Makris, T. M., Vu, V. V., Meier, K. K., Komor, A. J., Rivard, B. S., Münck, E., Que, L, Jr., and Lipscomb, J. D. “An Unusual Peroxo Intermediate of the Arylamine Oxygenase of the Chloramphenicol Biosynthetic Pathway”, J. Am. Chem. Soc., 137, 1608-1617 (2015) PMC
Knoot, C. J., Purpero, V. M., Lipscomb, J. D. “Crystal Structures of Alkylperoxo and Anhydride Intermediates in an Intradiol Ring-cleaving Dioxygenase” Proc. Natl. Acad. Sci., 112, 388-393 (2015) PMC
Lipscomb, J. D. “Life in a Sea of Oxygen” J. Biol. Chem. 289, 15141-15153 (2014) PMC
Makris, T. M., Knoot, C. J., Wilmot, C. M. and Lipscomb, J. D. “Structure of a Dinuclear Iron Cluster-Containing Beta Hydroxylase Active in Antibiotic Biosynthesis” Biochemistry, 52, 6662-6671 (2013) PMC
Aukema, K. G., Makris, T. M., Stoian, S. A., Richman, J. E., Münck, E., Lipscomb, J. D., and Wackett L. P. “Cyanobacterial aldehyde deformylase oxygenation of fatty aldehydes yields n-1 aldehydes and alcohols in addition to fatty alkanes”, ACS Catalysis, 3, 2228–2238 (2013) PMC
Banerjee, R., Meier, K. K., Münck, E., and Lipscomb, J. D. “Intermediate P* from Soluble Methane Monooxygenase Contains a Diferrous Cluster” Biochemistry, 52, 4331-4342 (2013) (Highlighted publication) PMC
Kovaleva, E. and Lipscomb, J. D. “Structural Basis for the Role of Tyrosine 257 of Homoprotocatechuate 2,3-Dioxygenase in Substrate and Oxygen Activation” Biochemistry, 51, 8755-8763 (2012) PMC
Mbughuni, M. M., Meier, K. K., Münck, E. and Lipscomb, J. D. “Substrate-Mediated Oxygen Activation by Homoprotocatechuate 2,3-Dioxygenase: Intermediates Formed by a Tyrosine 257 Variant” Biochemistry, 51, 8743-8754 (2012) PMC
Thompson, J. W., Salahudeen, A. A., Chollangi, S., Ruiz, J. C., Brautigam, C. A., Makris, T. M., Lipscomb, J. D., Tomchick, D. R., and Bruick, R. K. “Structural and Molecular Characterization of Iron-sensing Hemerythrin-like Domain within F-box and Leucine-rich Repeat Protein 5 (FBXL5)” J. Biol. Chem. 287, 7357-7365 (2012) (Cover illustration, paper of the year) PMC