Research Interests
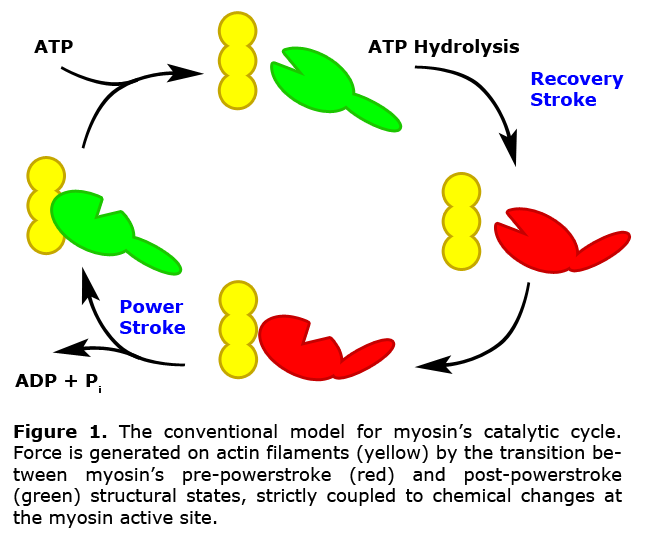
The generation of force in muscle contraction, while complex and nuanced in its full consideration, is nonetheless the direct result of a single interaction between two proteins, actin and myosin. This interaction represents the crucial device responsible for the shortening of muscle fibers, and in that partnership myosin takes the active role, unlocking the energy stored in ATP to pull on actin filaments. While a broad understanding of this force-generating cycle has been established for many years (Fig. 1), recent evidence suggests that myosin does not follow a progression of discreet structural transitions throughout the binding and hydrolysis of ATP, but rather constantly entertains a delicate balance of structural states within a population, achieving its function through subtle shifts in conformational equilibria.
The derivative of ATP bound to myosin at a given step in the hydrolytic cycle (myosin's "biochemical state") has long been associated with a particular protein conformation (myosin's "structural state"), but our lab has shown through spectroscopic analysis that populations with homogeneous biochemical states consistently display marked structural heterogeneity. This finding has important implications not only for the challenge of describing myosin's normal behavior, but also for understanding the fundamental nature of various disease phenotypes. My research focuses on providing new insight into the structural dynamics of myosin, using spectroscopy to characterize how distributions of structural states are affected by different physiological conditions.
Spectroscopic Investigation of Oxidative Stress in Myosin
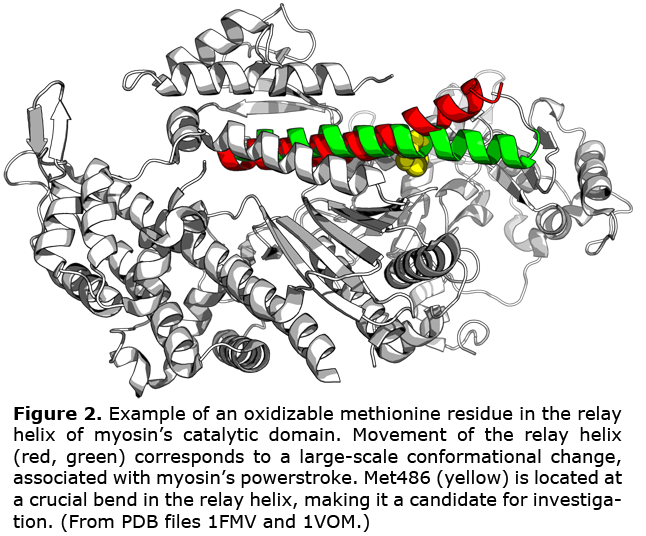
Reactive oxygen species (ROS) are produced as by-products of cellular respiration, and normally exist as essential components in a carefully-controlled redox environment. However, if the balance is preturbed and the proliferation of ROS overwhelms cellular antioxidant systems, the resulting oxidative conditions can lead to unchecked structural modification of proteins. We hypothesize that oxidation of myosin at various functionally-relevant sites contributes to the phenotypes observed in heart failure, muscle degeneration and biological aging, where elevated levels of ROS have also been observed (Fig. 2). Our lab has correlated ROS exposure with a decrease in myosin's ability to generate force both in vitro and in vivo. I seek to expand upon these observations by identifying specific residues on the myosin catalytic domain that impair function when oxidized, and using spectroscopy to observe the effects of site-specific oxidation on structural dynamics.